
Immune Response of Plants – Part 1
19.09.2017 / Scienceandmore / Category: Plant Biology
Plants represent a rich source of nutrients that is desired by microorganisms, which are mainly represented by bacteria but also fungi. To prevent exploitation, plants have developed an array of structural and chemical defence response mechanisms against infestation with pathogens. These pathogens are divided into the categories biotrophic, necrotrophic and hemibiotrophic. Biotrophic pathogens keep the host plants alive while feeding on them. Necrotrophic pathogens produce degrading enzymes to release plant nutrients and, thereby kill the plant. Hemibiotrophs behave in their early stages of infection like biotrophs, but eventually become necrotrophic during latter stages (1).
Structural plant defence
Their first line of plant defence is the structural defence and comprises of the waxy cuticle, consisting of a complex polymer of esterified fatty acids coated with waxes, and rigid cell walls as exterior surface that prevent pathogens from invading the plant. The structural defence is always present and does not need to be induced. This alone, however, is not adequate against all pathogens, so plants possess inducible immune responses that are activated upon pathogen attack (2,3).
Inducible plant defence
The currently well-established model of plants’ inducible immunity was proposed by Jones and Dangl in 2006 (4) and explains the interaction of plant and pathogen in a zig zag model. This model describes the possible interactions between plants and pathogens as two-pathed and four-phased, as follows. The first path consists of receptors on the surface of cells (pattern recognition receptors, abbreviated with PRRs) that recognise certain features of microorganisms that are called microbe-associated molecular patterns (MAMPs). This path is called pattern-triggered immunity (PTI), since the recognition of MAMPs can induce the plants immune response.
The second path is located mostly inside the cell and consists of certain plant proteins that are encoded by plant resistance genes (R genes). These plant proteins interact with proteins from the pathogenic microorganism that are called effectors or avirulence factors (encoded by avirulence genes) and aim to interfere with the plant immune response in order to alter/suppress it and mask the pathogen’s presence. This path is called effector-triggered immunity (ETI). Here, the term avirulence is somewhat misleading as it implies the loss of virulence, i.e. the loss of the ability to infect plants. Effectors (avirulence-factors) contribute to the pathogens virulence; however, these factors can be recognised by the plant proteins that are encoded by R genes. In case of recognition, plant immune responses are activated and the pathogen does not infect the plant and is, therefore, avirulent.
The four phases
In the first phase of plant-pathogen interaction, the plant recognises MAMPs and activates PTI to stop pathogen growth and colonisation. In the second phase, successful pathogens use effectors that interfere with the plant’s PTI, leading to effector-triggered susceptibility (ETS) in the plant. These effectors thus increase the pathogen’s virulence. If the plant recognises the microbial effectors with R gene-encoded proteins, then the third phase is triggered, activating ETI. In the fourth phase, natural selection favours the pathogens which have abandoned or modified their recognized effectors, or alternatively gained unrecognised effectors, meaning that the pathogens suppress ETI, again leading to ETS (see figure) (4). It is important to understand that this is not a chronological sequence where PTI is followed by ETS, followed by ETI, and so on, but should rather be considered as evolutionary adjustments to increase virulence on the pathogen side (abandonment/modification of recognized effectors, or gain of unrecognised effectors) and immunity on the plant side (recognition of MAMPs and effectors). It could be compared to an “arms race” on an evolutionary timescale.
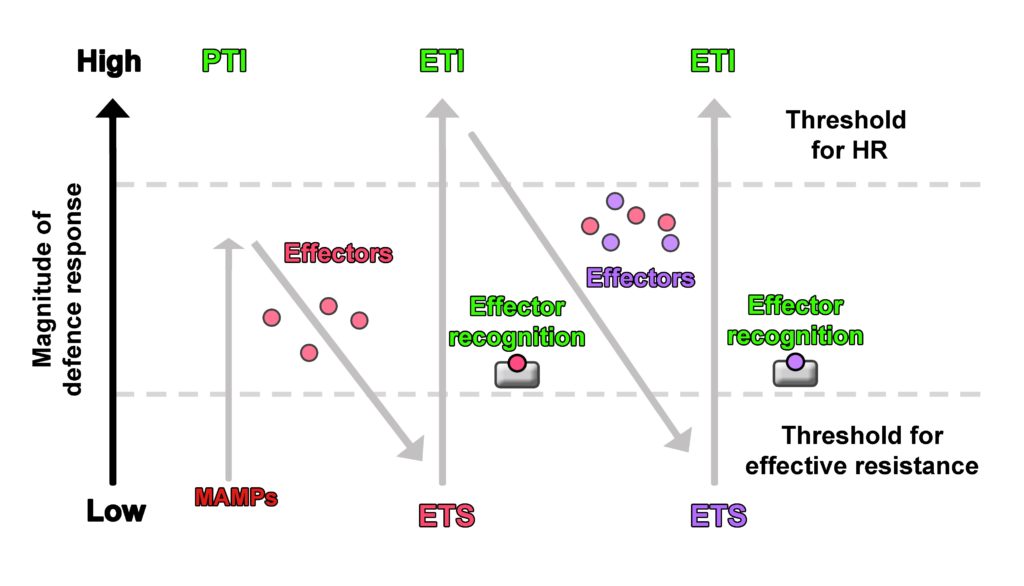
Zig Zag model of plant immune response against pathogens. HR = Hypersensitive Response (in reference to Jones and Dangl in 2006).
Pattern-triggered immunity (PTI)
Pathogenic bacteria can enter through stomata (surface openings that are needed for gas exchange of the plant) and other openings such as wounds, into the plant where they proliferate. Fungi enter directly into epidermal cells or penetrate the plant with their hyphae (filamentous structures of fungi) between and through cells. At this point, the plant’s PRRs perceive MAMPs and transmit signals to the cell (4-6).
The best described MAMPs are fungal chitin, bacterial peptidoglycans and lipo-polysaccharides, and bacterial flagellin (1,6,7). MAMPs are in general abundance and their nature is essentiality for microorganisms. Fungal chitin is an essential integral component of fungal cell walls and estimated to be the second most abundant polysaccharide in the world (besides cellulose); bacterial peptidoglycans are essential integral components of bacterial cell walls; lipopolysaccharides are membrane components of Gram-negative bacteria; bacterial flagellin is a component of the bacterial flagellum, which is used for locomotion (widespread among different species). This essential character often requires conservation throughout evolution and alteration could lead to a loss of microbial fitness and virulence for pathogens. The flagellum for example is not essential for bacterial survival, but contributes strongly to the virulence of bacterial pathogens (5).
Activation of PTI characteristically triggers reactive oxygen species (ROS) production, expression of pathogen-resistance genes (PR genes), callose deposition in the plant´s cell walls and closure of stomata as the main reactions to adjust the plant to pathogen attacks (1,4,7). ROS act as signalling molecules inside the plant but also as defence components, due to their reactive properties, to damage and kill pathogens. PR gene products can be active components against pathogens or can be involved in signal transmission. Callose strengthens the cell wall, and the closure of stomata decreases possible entry points for additional pathogens.
Effector-triggered immunity (ETI)
As stated before, pathogens additionally use effectors to alter the plants immune response, and mask their presence from the plant. Effectors increase the plant’s susceptibility, enhance microbial fitness, and can cause nutrient leakage for nutritional purposes of the pathogen (4,5,9).
At this point, microbial effectors may be either directly or indirectly recognised by the plant’s R-gene products, which leads to an activation of ETI. There are two hypotheses that describe the recognition of microbial effectors by plant’s R gene products. The first states that the R gene products in general directly recognise microbial effectors, and this is called the gene-for-gene hypothesis. The second, called the guard hypothesis, describes indirect effector recognition by R gene products. Effectors manipulate certain plant components as their targets and, rather than being recognised directly, R gene products recognise the resultant alteration of the targeted components. Therefore, R gene products ‘guard’ specific components and recognise a ‘modified-self’. This way, a limited set of R gene products can cover a wider range of microbial effectors. Most R genes encode NB-LRR (nucleotide-binding leucine-rich repeat) proteins, and only approximately 150 NB-LRR genes are found in A. thaliana, an important model plant in biology; a number that would be insufficient to cover all known and potential microbial effectors. This fact supports the guard hypothesis (3,4,9,10).
PTI is understood to be the first line of defence against the majority of pathogens, mainly triggering weaker and rather localised defence responses, such as ROS accumulation, while ETI is more specific and triggers mainly stronger responses, such as hypersensitive response (HR). HR is a form of programmed cell death that is triggered at the site of infection and is understood to cut off pathogens from water and nutrients, thereby limiting proliferation (9,12). However, the reversed case is not excluded and suggests that the specific character of the MAMP or effector, as well as their quantity and exposure time, together modulate a specific plant immune response (8,11).
Recently, the zig zag model for plant-pathogen interaction was criticised since it is based on the plant’s interaction with biotrophic microbes, i.e. pathogens that keep their host plans alive while feeding on them, and lacked integration of symbiotic interactions, or the response to necrotrophic pathogens where HR would rather be a favourable result, since necrotrophic pathogens aim to kill plant cells in order to obtain nutrients. HR, which is a form of programmed cell death would, therefore, contribute to an invasion by necrotrophs and be counterproductive for the plant.
References
1. Freeman, B. C. and Beattie, G.A. (2008) An Overview of Plant Defenses against Pathogens and Herbivores. The Plant Health Instructor. Available from: DOI: 10.1094/PHI-I-2008-0226-01
2. Sels, J., Mathys, J., De Coninck, B.M.A., Cammue, B.P.A., and De Bolle, M.F.C. (2008) Plant pathogenesis-related (PR) proteins: A focus on PR peptides. Plant Physiology and Biochemistry 46:941-50.
3. Kim, M.G., da Cunha, L., McFall, A.J., Belkhadir, Y., DebRoy, S., Dangl, J.L., Mackey, D. (2005) Two Pseudomonas syringae Type III Effectors Inhibit RIN4-Regulated Basal Defense in Arabidopsis. Cell, 121(5), 749-59.
4. Jones, J.D. and Dangl, J.L. (2006) The plant immune system. Nature, 444, 323-9.
5. Pel, M.J.C. and Pieterse, C.M.J. (2013) Microbial recognition and evasion of host immunity. Journal of Experimental Botany, 64(5), 1237-48.
6. Yeh, Y.H., Chang, Y.H., Huang, P.Y., Huang, J.B., Zimmerli, L. (2015) Enhanced Arabidopsis pattern-triggered immunity by overexpression of cysteine-rich receptor-like kinases. Front. Plant Sci., 6, 322.
7. Quirino, B. F. and Brent, A. F. (2003) Deciphering host resistance and pathogen virulence: the Arabidopsis/ Pseudomonas interaction as a model. Molecular plant pathology, 4(6), 517-30.
8. Doehlemann, G. and Hemetsberger C. (2013) Apoplastic immunity and its suppression by filamentous plant pathogens. New Phytologist, 198, 1001-16.
9. Coll, N.S., Epple, P., Dangl, J.L. (2011): Programmed cell death in the plant immune system. Cell Death and Differentiation, 18, 1247-56.
10. Alfano, J. R., Collmer, A. (2004) TYPE III SECRETION SYSTEM EFFECTOR PROTEINS: Double Agents in Bacterial Disease and Plant Defence. Annual Review of Phytopathology, 42, 385-414.
11. Nishimura, M.T. and Dangl, J.L. (2010) Arabidopis and the plant immune system. The Plant Journal. 61, 1053-1066.
12. Torres, M.A., Jones, J.D.G., Dangl, J.L. (2006) Reactive Oxygen Species Signaling in Response to Pathogens. Plant Physiol., 141(2), 373-8.
